Human health and disease emerge from complex interactions across molecular, cellular, and population scales.
We use AI to integrate and analyse heterogeneous data across all layers of health - from genomics and medical imaging to clinical records and population studies - to develop interpretable and robust models that support medical decision-making. Building on expertise in machine learning and biomedicine, we strive to improve health outcomes through earlier diagnosis, better risk assessment, and more precise prevention and treatment.
Life is complex, and complicated: living systems are composed of millions of different molecular entities, interacting in space and time to create emergent properties.
We use AI to design and execute sophisticated and large-scale experiments and studies using modern biotechnologies and instruments; organise, navigate and reason with massive amounts of facts and data; and create interpretable, predictive, scientifically founded models of biological systems at scales from nano- to Megameters.
Going to smaller and smaller distances, the physical laws behind Life turn into the fundamental laws behind chemistry and physics.
From a scientific AI perspective, the collisions of elementary particles, the astrophysical evolution of our Universe, the dynamics of molecules and cells, and the collective behavior of tissues, organisms, and even ecosystems can be understood with similar numerical methods. Our unifying goal is to understand fundamental structures and principles across scales from complex, high-dimensional data. We develop AI-methods to uncover the underlying fundamental mechanisms by combining theory, simulation, and optimal analysis of large-scale experimental and observational data. The requirements of the different research fields in terms of accuracy, uncertainty quantification, robustness, and interpretability challenge our methods and tools and drive our interdisciplinary program on AI in the sciences.
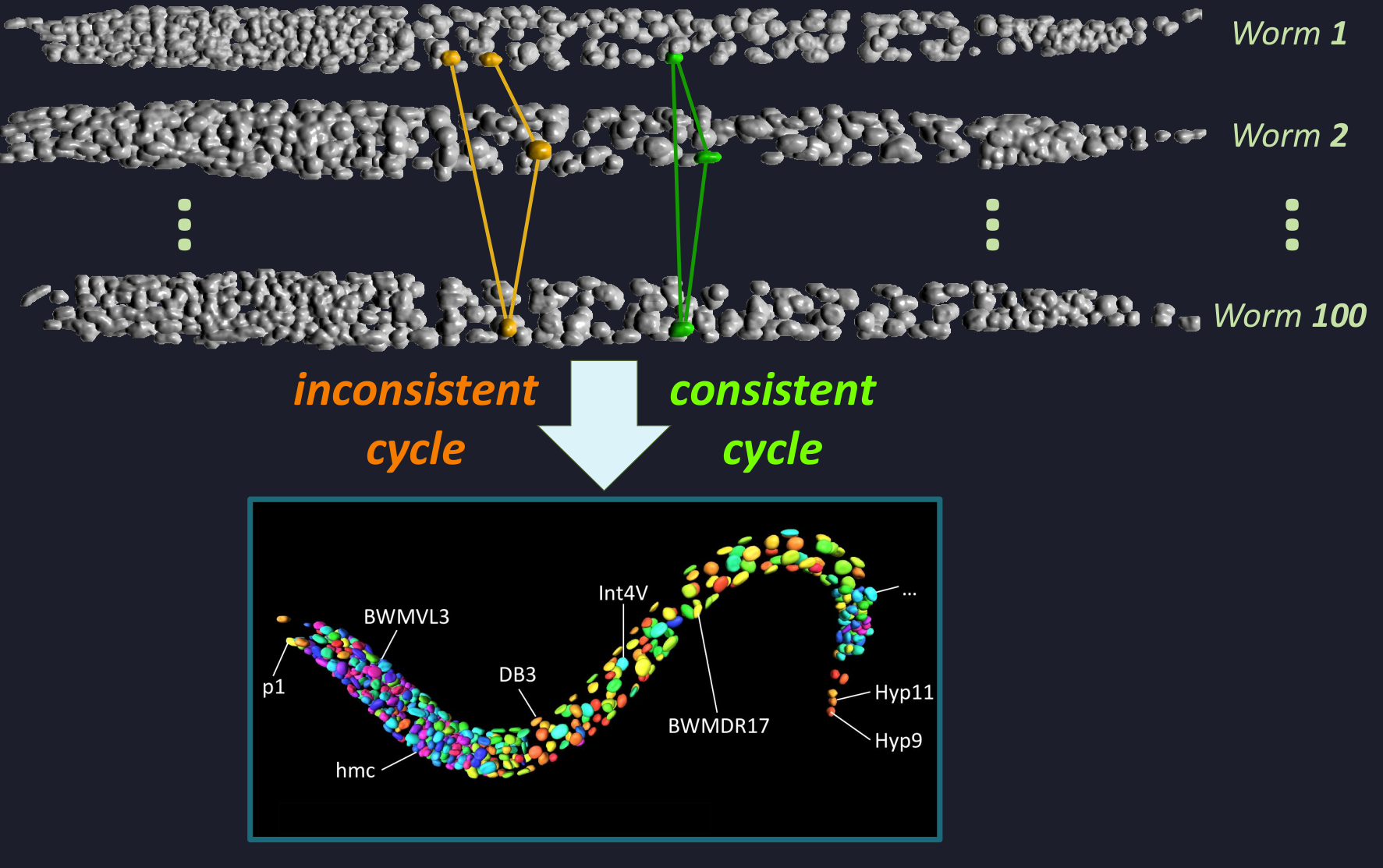
Stereotypical organisms share the same cellular body plan across individuals, enabling biological observations from different specimens to be aligned cell by cell. In the model organism Caenorhabditis elegans, fluorescence microscopy can measure gene expression across its cells, but constructing a unified cellular atlas requires establishing consistent cell correspondences across many specimens—a task that is extremely challenging to perform manually at scale.
We develop WormMatch, an AI-driven framework for unsupervised atlas discovery that formulates this problem as large-scale multi-matching. By combining machine learning with combinatorial optimization, WormMatch jointly aligns cells across hundreds of microscopy images while learning matching structure directly from data without requiring extensive annotations.
This approach enables automated construction of cellular reference atlases and scalable integration of gene-expression measurements, providing a new computational foundation for studying biological programs encoded in the genome.
Applying state-of-the-art segmentation models to new medical imaging datasets typically requires extensive manual optimization and expert knowledge. We develop adaptive and interactive AI systems that remove this barrier — from fully automated configuration to real-time and language-guided interaction.
nnU-Net is a self-configuring segmentation framework that automatically adapts to new datasets and delivers out-of-the-box 2D and 3D models, making state-of-the-art medical image segmentation broadly accessible (8,000 citations and 300+ daily downloads).
nnDetection extends this automation to 3D object detection, self-configuring for arbitrary volumetric tasks and achieving state-of-the-art performance without manual intervention.
nnInteractive enables expert-guided 3D segmentation within tools such as Napari and MITK, translating intuitive prompts into accurate volumetric masks across 120+ datasets.
VoxTell brings free-text–guided universal 3D segmentation, mapping clinical language directly to volumetric masks and generalizing across modalities and unseen concepts.
Medical imaging AI research is largely driven by data availability rather than clinical need, leading to benchmarks that may reward performance on narrow, repetitive tasks with limited real-world impact. As a result, current benchmarks often provide only partial insight into whether emerging medical imaging AGI systems can address the most impactful clinical problems. We introduce MEDAL (Medical Imaging AGI’s Last Exam), a next-generation benchmark designed to assess the true clinical capabilities of medical imaging AGIs. MEDAL shifts benchmarking toward a clinically driven, top-down approach in which high-impact medical imaging problems are defined by a global expert community. These problems are paired with diverse, previously unpublished multimodal datasets crowdsourced worldwide, with data and task contributions incentivized through a total reward of 1 million EUR. Together with a statistically grounded evaluation framework, MEDAL enables reliable assessment of generalization, clinical relevance, and transformative potential, establishing a new gold standard for medical imaging AGI evaluation.
NAILIt develops new approaches that will allow future AI systems to adapt to changing conditions – such as new tasks or unexpected situations – with the flexibility and versatility known from living organisms. At the core of NAILIt lies the question of how the learning principles observed in animal brains can be transferred to AI. Whereas modern AI models – such as large language models – are typically trained once on massive datasets and then operate with fixed parameters, animals continually adjust their behavior to new situations. They do so rapidly, efficiently, and with minimal effort. Such adaptive capabilities are becoming increasingly important for AI systems used in real-world scenarios, for example in autonomous vehicles or in interactive AI agents that engage directly with humans. Self-developed AI tools are used for dynamical systems reconstruction (DSR) to derive generative models of learning from neural and behavioural data. In the long term, the developed methods will also be used in psychiatric contexts, for example to predict individual disease progression or to control adaptive neurofeedback procedures.
MadGraph is one of a few simulation tools providing the theory predictions for the multi-purpose LHC experiments. It starts from a Lagrangian and uses Quantum Field Theory to generate LHC scattering processes, compute transition amplitudes, describe QCD jet radiation, hadronize the QCD partons, decay hadrons, and predict all particles entering the LHC detectors. Based on a freely chosen world of elementary particles and their interaction, these simulated events will then be compared to observed LHC events, to infer the correct theory. The upcoming high-luminosity LHC (2028-2041) needs simulations with significantly improved precision and speed. Classically a Monte Carlo simulation, the next MadGraph release will benefit from modern machine learning in all possible ways, from end-to-end ML-generators to neural multi-channel importance sampling, learned phase space mappings, precision amplitudes encoded in uncertainty-aware surrogates, and ML-adapted tools to evaluate higher-order QCD corrections, to reweighted fragmentation functions (MLhad), all supported by MadAgents.
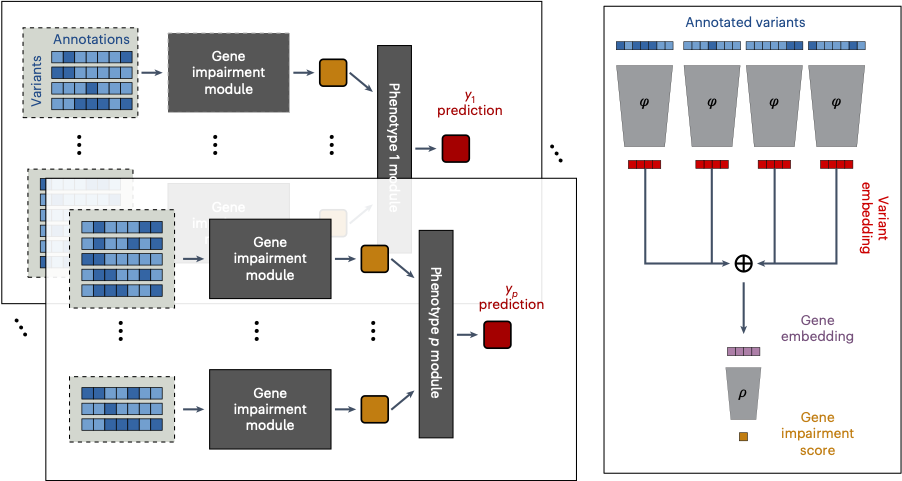
Rare genetic variants can have large effects on human health, but their impacts are difficult to detect with standard methods. We developed DeepRVAT, a rare variant analysis framework for modeling disease risk and biological traits through the integration of rare variant annotations.
Using deep set networks, DeepRVAT learns gene-level impairment scores from variant annotations (e.g. conservation and splicing-related scores), boosting power for association testing in large biobank datasets. Compared with existing approaches, it increases gene discovery and improves identification of individuals with high genetic risk in population-scale studies - supporting deeper insights into disease biology and more informative genetic risk stratification.
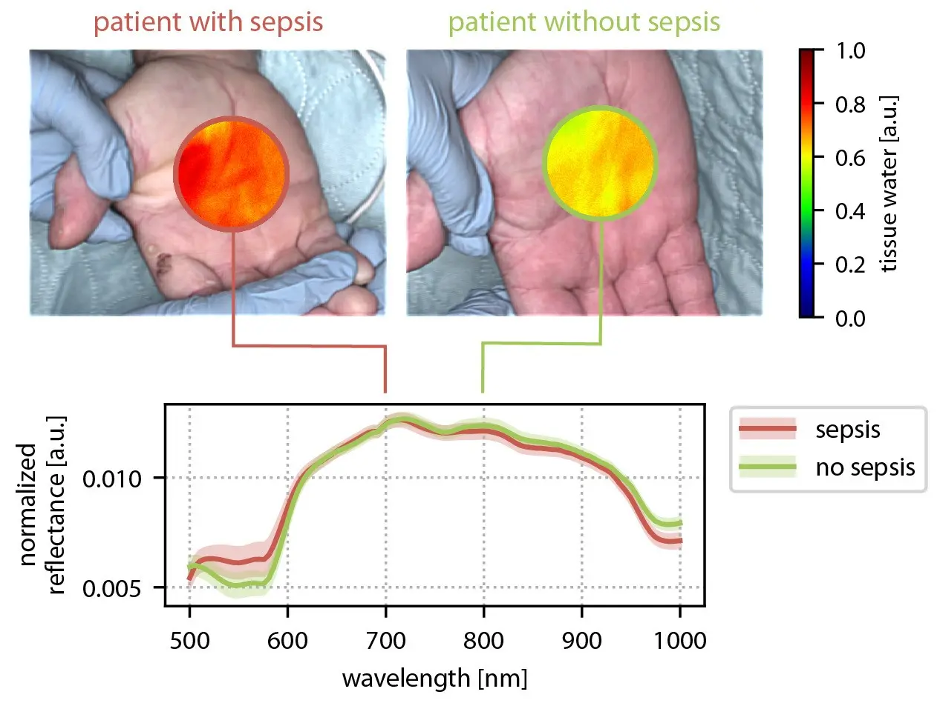
Spectral imaging methods such as multispectral diffuse reflectance imaging and photoacoustic imaging can non-invasively reveal functional tissue properties, but robust quantification in clinical practice remains challenging - largely due to the inverse nature of the reconstruction problem and the scarcity of labeled reference data. Neural Spicing reframes this challenge as a decoding task addressed with physics-constrained neural networks. A core innovation is a learning-to-simulate approach that leverages unlabeled and weakly labeled measurements to make simulated spectral images more realistic, narrowing the domain gap and improving quantification accuracy while explicitly handling uncertainty. The long-term vision is a second generation of safe, low-cost spectral imaging that can translate into impactful applications from interventional monitoring to improved diagnosis and therapy planning across multiple diseases. First clinical success stories include non-invasive sepsis diagnosis and contrast agent-free, real-time ischemia monitoring in laparoscopic surgery.
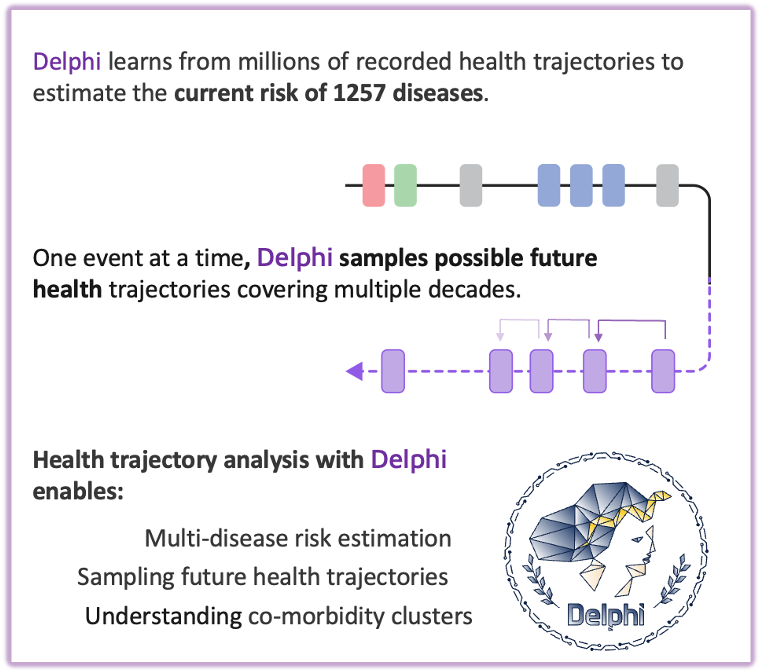
Many chronic diseases develop gradually, shaped by complex interactions between prior diagnoses, lifestyle factors, and the timing of medical events. Delphi-2M addresses this challenge by modeling long-term risk trajectories across more than 1,000 conditions, forecasting health outcomes up to ten years in advance.Inspired by concepts from large language models, the framework treats medical histories as ordered sequences of events and learns patterns in disease progression over time. Trained on anonymised data from 400,000 UK Biobank participants and validated in 1.9 million individuals from the Danish National Patient Registry, Delphi-2M shows strong performance for conditions with consistent progression patterns, including selected cancers and cardiovascular diseases.Developed through a collaboration between EMBL, DKFZ, the Universities of Copenhagen and Helsinki, the project lays the groundwork for data-driven preventive health research and scalable, personalised risk modeling.
ORena SAVE FOCUS is the first challenge to test whether AI can truly understand full-length surgeries in order to solve crucial clinical problems that matter everywhere in the world.
The Medical Goal: To help prevent retained surgical items by enabling models to track, contextualize, count, and localize foreign objects, such as sponges and needles, throughout an entire surgical procedure.
The Technical Goal: To benchmark and advance long-context video understanding, spanning the full spectrum from single-frame perception to procedure-level surgical understanding.

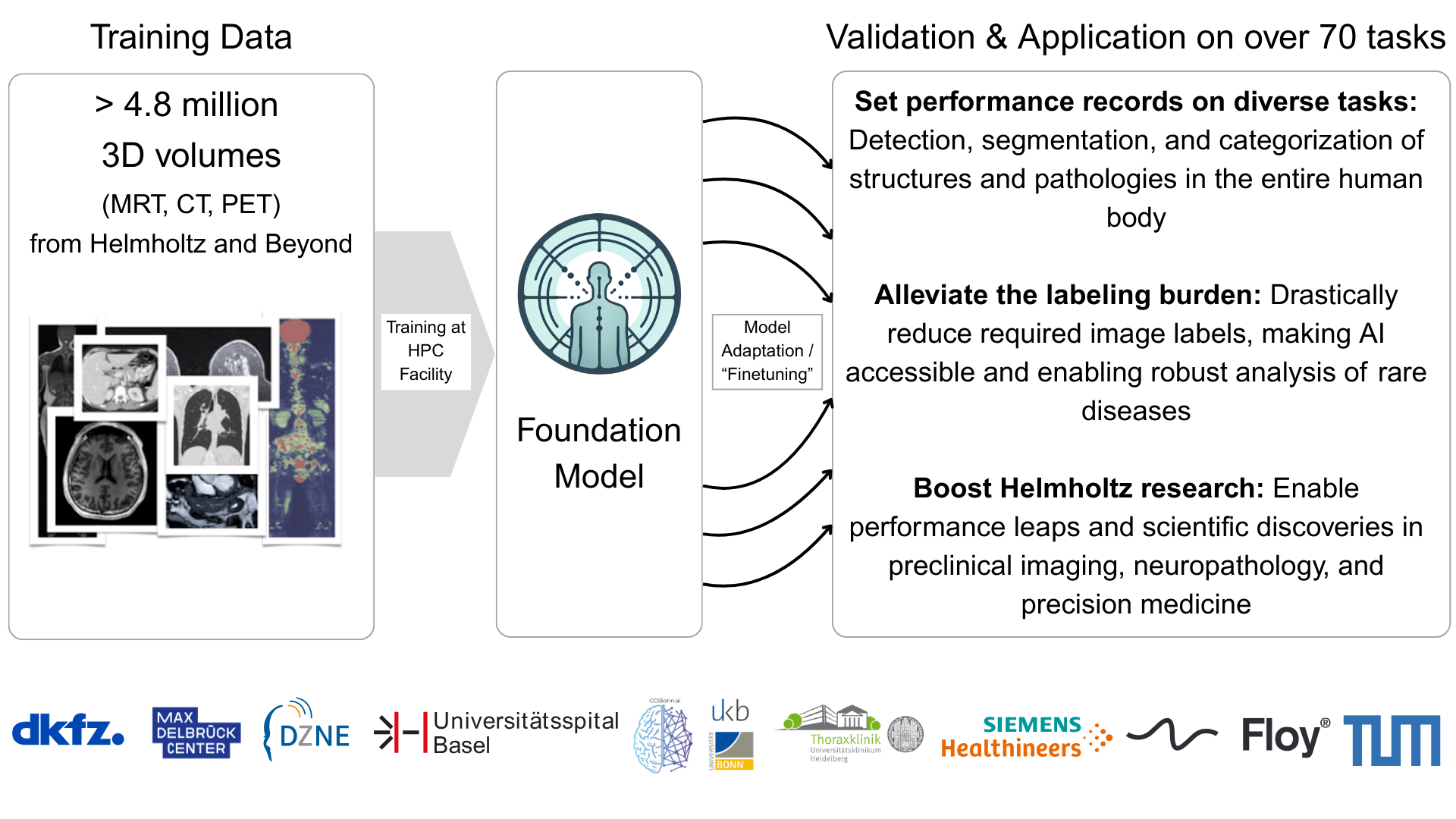
The Human Radiome Project (THRP) develops an AI-powered foundation model to unlock the full complexity of 3D radiological imaging. Funded by the Helmholtz Association’s Foundation Model Initiative (HFMI), THRP aims to build a digital framework that integrates structural, functional, and pathological imaging information.
THRP trains a self-supervised foundation model on over 3 million 3D image volumes (including CT and MRI). The dataset combines clinical repositories at DKFZ and partner hospitals in Bonn, Heidelberg, and Basel with population studies and public datasets, enabling robust, generalizable representations.
As medical imaging demand grows and clinical expertise becomes scarce, THRP enables scalable AI for diverse radiology tasks without extensive task-specific annotation. Long term, it aims to accelerate research and advance personalized healthcare by integrating imaging with other data modalities.